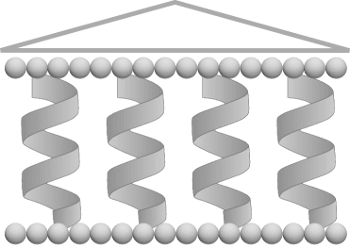

The TMPL database1 contains 3D structures of transmembrane proteins with bilayer boundaries assigned by the ANVIL algorithm2 and represented by two planes.
The structural quality of the transmembrane domains delimited by ANVIL is evaluated with the MAIDEN statistical potential3; both methods are available through the OREMPRO web server4.
The molecular structures in TMPL are i) either atomistic or coarse-grained representations of proteins, and ii) either theoretical or experimental models.
The structural quality of the transmembrane domains delimited by ANVIL is evaluated with the MAIDEN statistical potential3; both methods are available through the OREMPRO web server4.
The molecular structures in TMPL are i) either atomistic or coarse-grained representations of proteins, and ii) either theoretical or experimental models.
1Postic G, Ghouzam Y, Etchebest C, Gelly JC. Database (2017)
doi:10.1093
2Postic G, Ghouzam Y, Guiraud V, Gelly JC. Protein Eng Des Sel (2016) 29:87-92
3Postic G, Ghouzam Y, Gelly JC. Biochimie (2015) 115:155-161
2Postic G, Ghouzam Y, Guiraud V, Gelly JC. Protein Eng Des Sel (2016) 29:87-92
3Postic G, Ghouzam Y, Gelly JC. Biochimie (2015) 115:155-161
Statistics
| • Native structures: 997 | • Predicted models: 10,170 |
| ⤷ Alpha-helical: 830 | • Pfam domains: 729 |
| ⤷ Beta-sheet: 167 | • Deposited models: 11 |
• Martinized: 1,039

Coarse-grained models compatible with the MARTINI force field (Marrink et al., 2007), and produced from native structures by using the martinize.py script (de Jong et al., 2013).
|
Users have the possibility of adding new entries to the TMPL database, by uploading protein structures orientated in the lipid bilayer. This can be achieved by using either available membrane protein structures (native or predicted), or new theoretical models, such as those that users can generate with the ORION web server5,6.
4Postic G, Ghouzam Y, Gelly JC. Bioinformatics (2016)
32:2548-2550
5Ghouzam Y, Postic G, de Brevern AG, Gelly JC. Bioinformatics (2015) 31:3782-3789
6Ghouzam Y, Postic G, Guerin PE, de Brevern AG, Gelly JC. Sci Rep (2016) 6:28268
5Ghouzam Y, Postic G, de Brevern AG, Gelly JC. Bioinformatics (2015) 31:3782-3789
6Ghouzam Y, Postic G, Guerin PE, de Brevern AG, Gelly JC. Sci Rep (2016) 6:28268